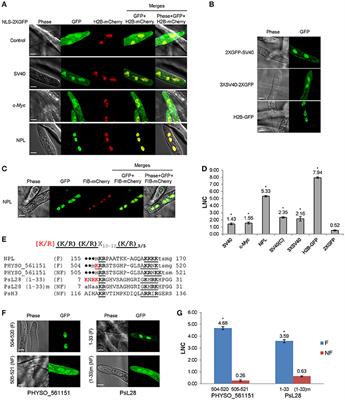
SeqNLS: Nuclear Localization Signal Prediction Based on Frequent Pattern Mining and Linear Motif Scoring | PLOS ONE
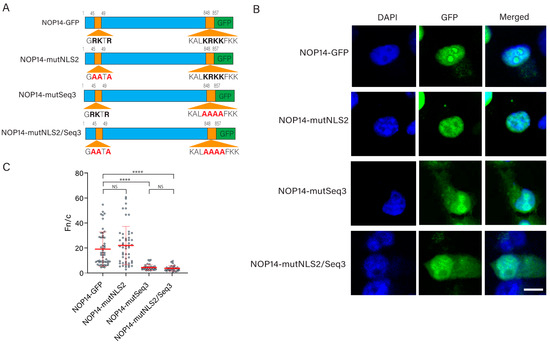
Cells | Free Full-Text | Bioinformatics and Functional Analysis of a New Nuclear Localization Sequence of the Influenza A Virus Nucleoprotein
SeqNLS: Nuclear Localization Signal Prediction Based on Frequent Pattern Mining and Linear Motif Scoring | PLOS ONE

Nuclear Localization Signal | In Silico Nuclear Localization Signal Prediction | NLS Mapper| - YouTube

Figure 2.3 from Nuclear export signals (NESs) in Arabidopsis thaliana : development and experimental validation of a prediction tool | Semantic Scholar
NES and NLS prediction in the bHLH domain of TCF4. (A) Results of NES... | Download Scientific Diagram

Figure 2.1 from Nuclear export signals (NESs) in Arabidopsis thaliana : development and experimental validation of a prediction tool | Semantic Scholar
SeqNLS: Nuclear Localization Signal Prediction Based on Frequent Pattern Mining and Linear Motif Scoring | PLOS ONE

LLRC, LLR, and NLS model fits for each 1D (a,b), and 2D (c) predictor... | Download Scientific Diagram

Prediction results for putative NLS, NES and DNA-binding motifs on CAV... | Download Scientific Diagram

Identification of a nuclear localization signal in the Plasmodium falciparum CTP: phosphocholine cytidylyltransferase enzyme | Scientific Reports

NLS Mapper, PredictProtein and COMPARTMENT OAS3 nuclear localization... | Download Scientific Diagram

DNA binding residues in region F1 acts as a Nuclear Localization Signal... | Download Scientific Diagram

Cells | Free Full-Text | Bioinformatics and Functional Analysis of a New Nuclear Localization Sequence of the Influenza A Virus Nucleoprotein

NLStradamus: a simple Hidden Markov Model for nuclear localization signal prediction | BMC Bioinformatics | Full Text

Discovering nuclear targeting signal sequence through protein language learning and multivariate analysis - ScienceDirect

Analysis of rice nuclear-localized seed-expressed proteins and their database (RSNP-DB) | Scientific Reports

Analysis of rice nuclear-localized seed-expressed proteins and their database (RSNP-DB) | Scientific Reports

Identification of a bipartite nuclear localization signal in the silkworm Masc protein - Sugano - 2016 - FEBS Letters - Wiley Online Library
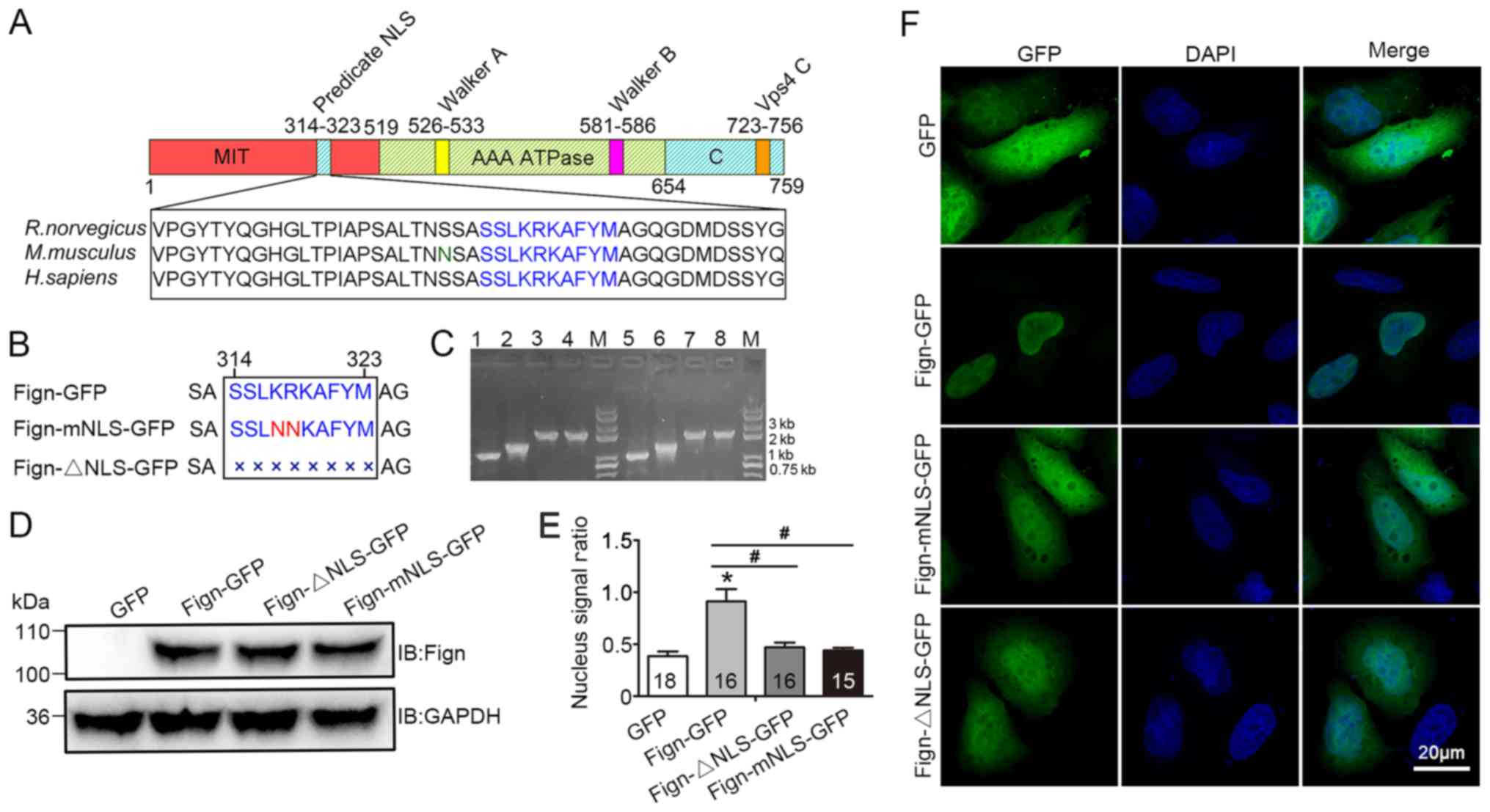
A nuclear localization signal is required for the nuclear translocation of Fign and its microtubule‑severing function

Cells | Free Full-Text | Bioinformatics and Functional Analysis of a New Nuclear Localization Sequence of the Influenza A Virus Nucleoprotein